Welcome to SIM-XL
State-of-the-art structural mass spectrometry-based proteomics is often accomplished by using cross-linkers to covalently bind two or more amino-acid groups. This strategy is complementary to classical structural biology and therefore broadens the toolset for analyzing protein and protein-protein complex structures.
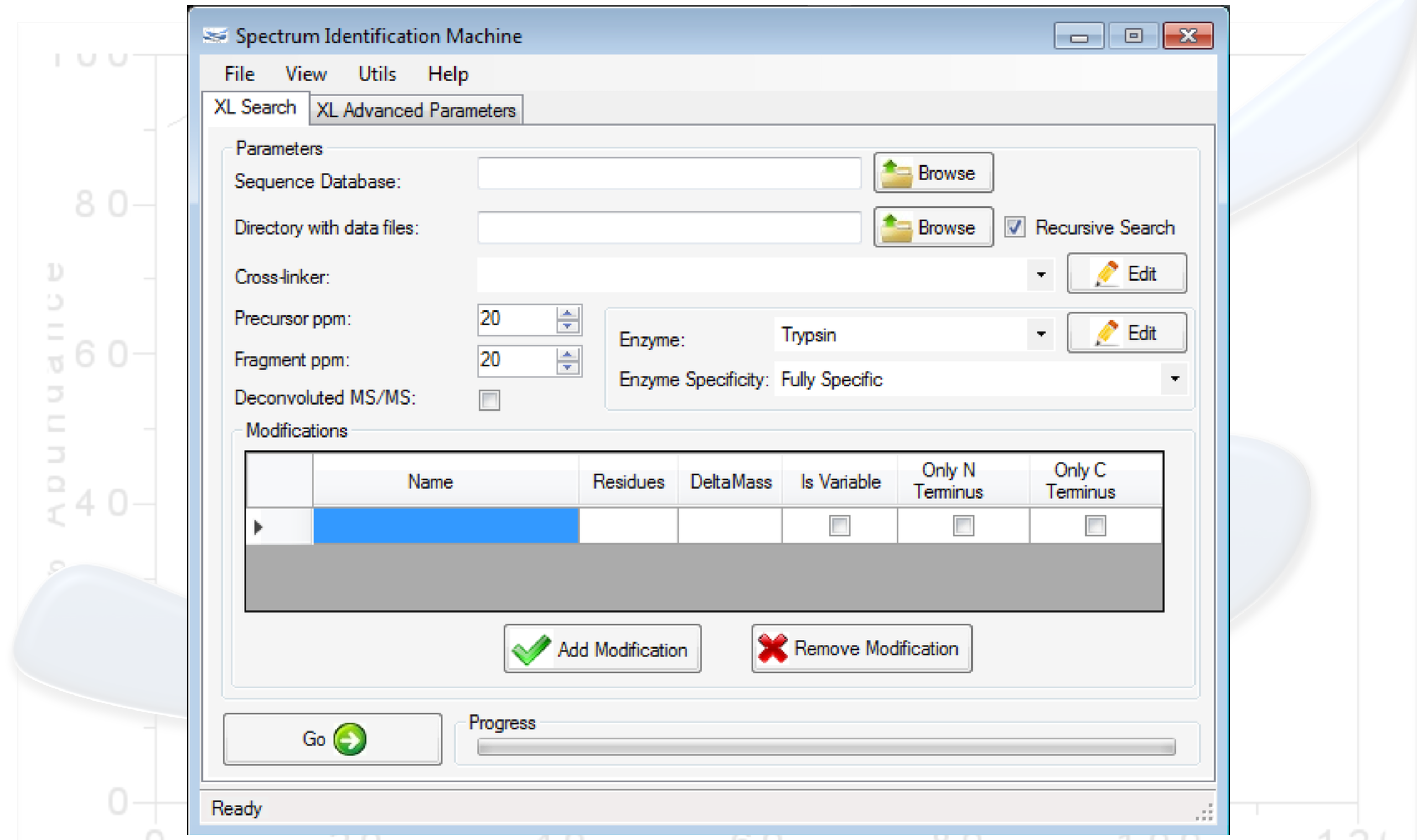
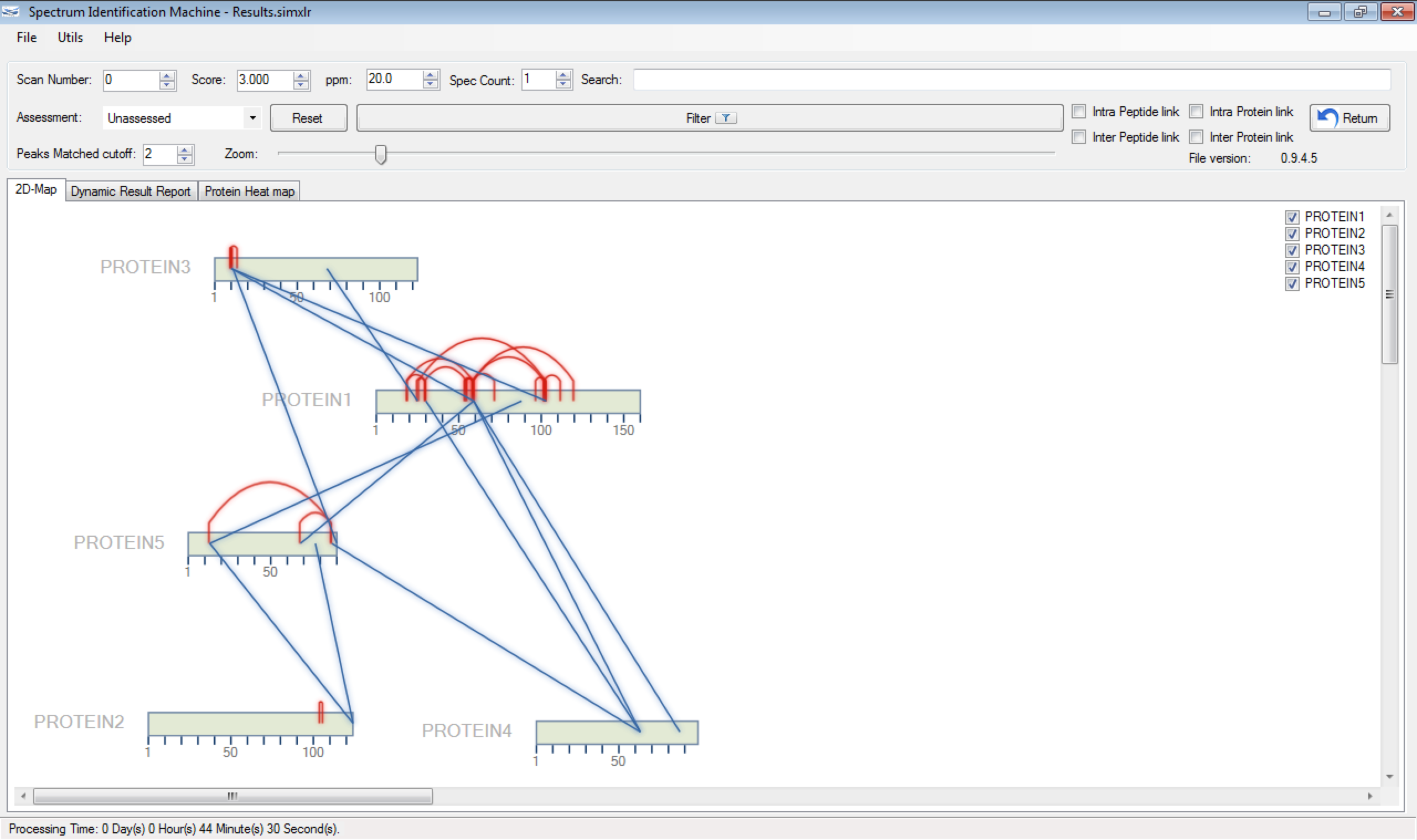
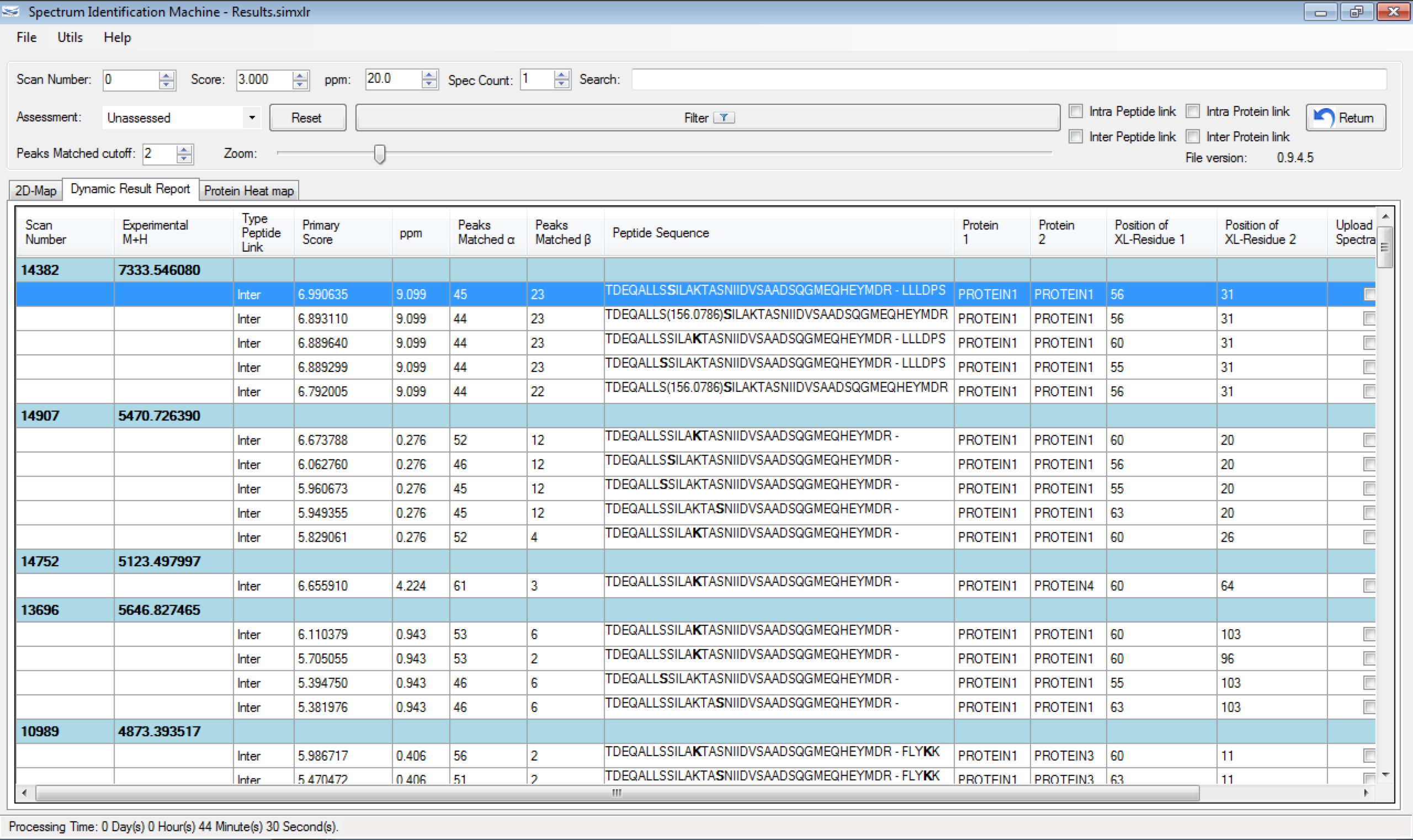
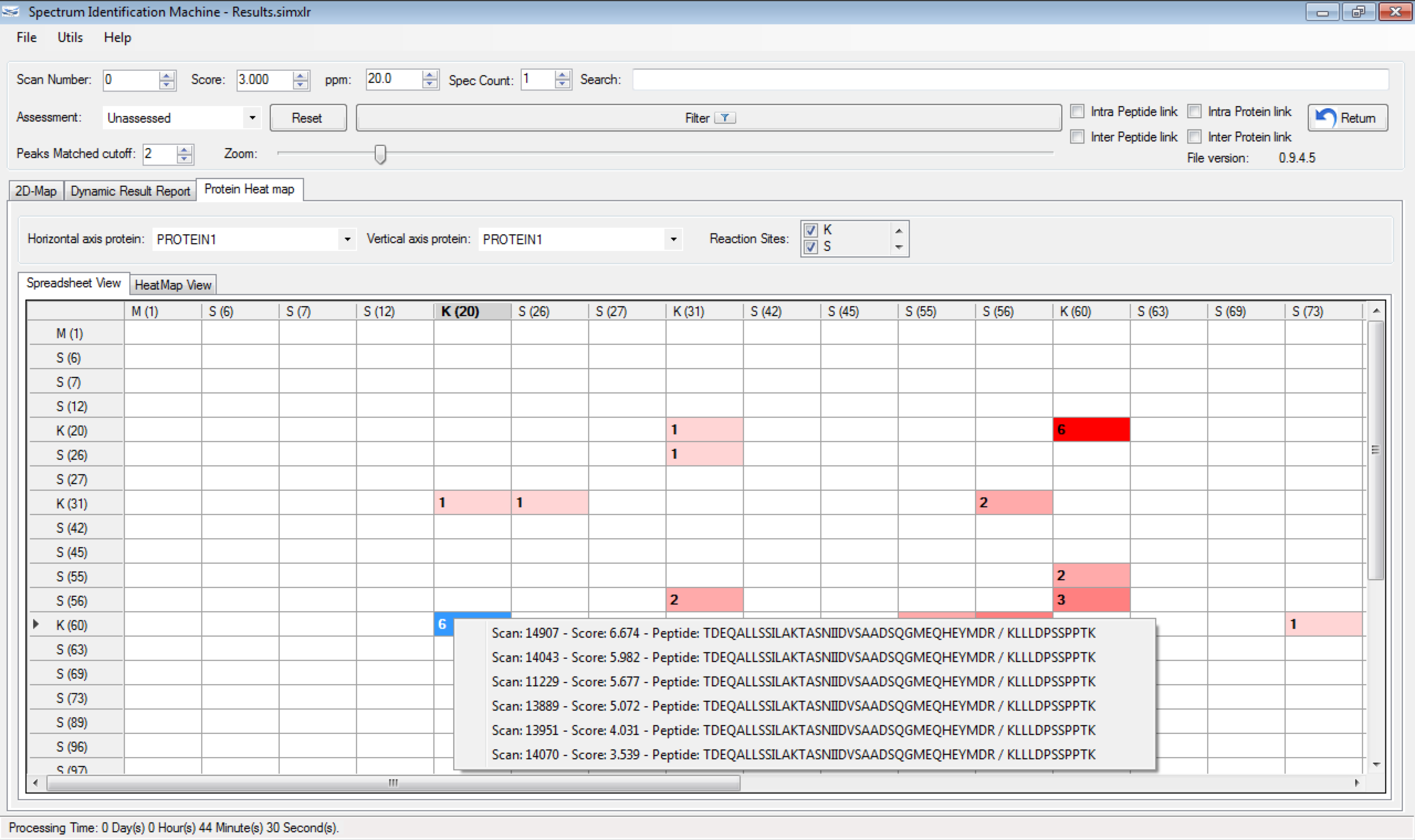
One of the greatest challenges in identifying cross-linked peptides in complex protein mixtures is computationally dealing with the large search space, which grows exponentially with each peptide included in the sequence database. Spectrum Identification Machine for Cross-Linked Peptides (SIM-XL) software uses an algorithm that overcomes this limitation by capitalizing on experimental features that allows it to effectively address the massive combinatorial problem at hand and presents the results in a user friendly manner. Thus, SIM-XL is recommended for studies dealing with protein structure and protein-protein interaction in either simple or complex protein mixtures.
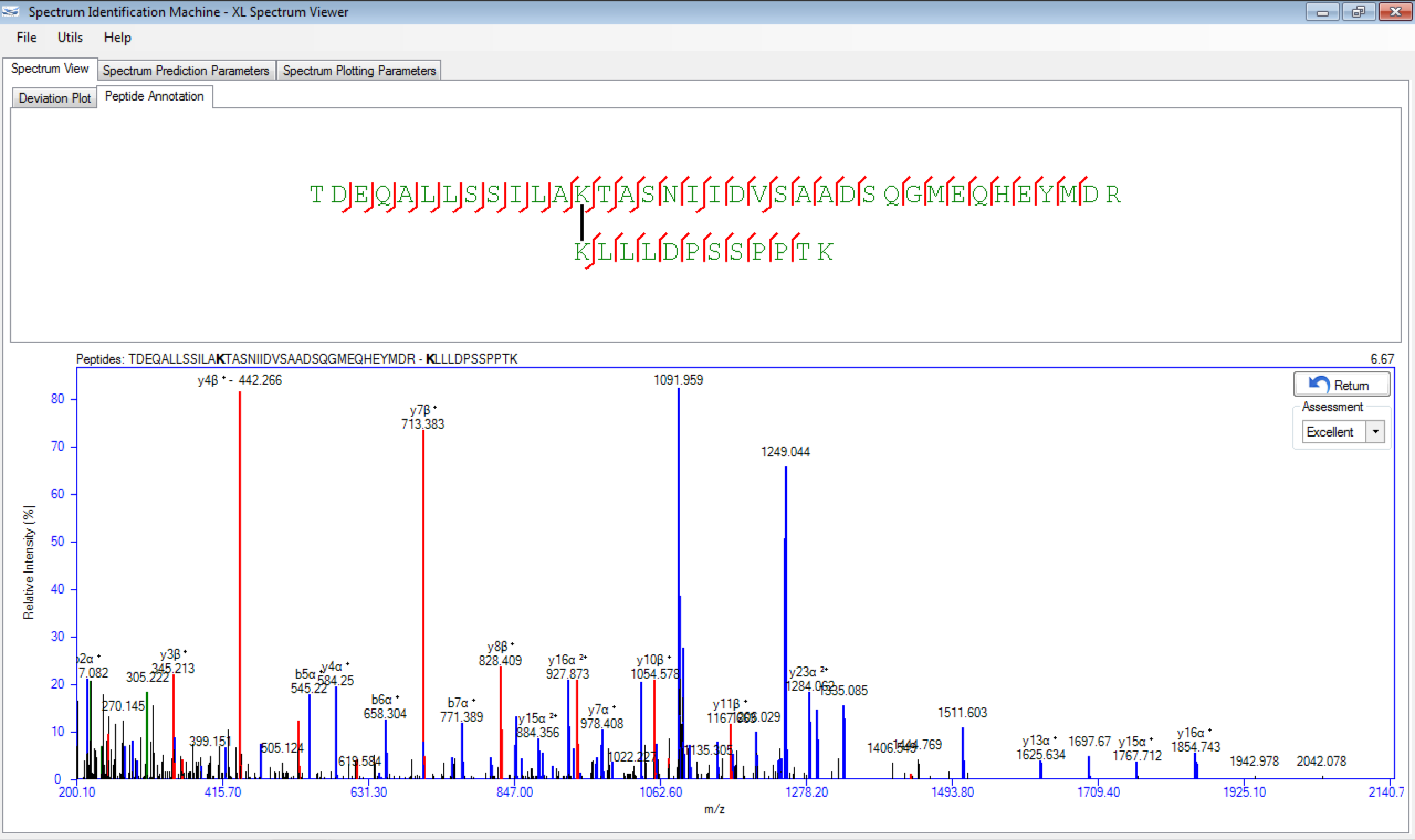
SIM-XL is user-friendly, simple and provides a graphical user interface.
View a video demonstrating SIM-XL in action!
 Diogo B Lima∗, John T Melchior∗, Jamie Morris, Valmir C Barbosa, Julia Chamot-Rooke, Mariana Fioramonte, Tatiana A C B Souza, Juliana S G Fischer, Fabio C Gozzo, Paulo C Carvalho, and W. Sean Davidson, “Characterization of homodimer interfaces with cross-linking mass spectrometry and isotopically labeled proteins”, Nature Protocols (13), pages 431–458, 2018.
Diogo B Lima∗, John T Melchior∗, Jamie Morris, Valmir C Barbosa, Julia Chamot-Rooke, Mariana Fioramonte, Tatiana A C B Souza, Juliana S G Fischer, Fabio C Gozzo, Paulo C Carvalho, and W. Sean Davidson, “Characterization of homodimer interfaces with cross-linking mass spectrometry and isotopically labeled proteins”, Nature Protocols (13), pages 431–458, 2018.
 Diogo B Lima, Tatiani B de Lima, Tiago S Balbuena, Ana Gisele C. Neves-Ferreira, Valmir C Barbosa, Fabio C Gozzo, Paulo C Carvalho, “SIM-XL: a powerful and user-friendly tool for peptide cross-linking analysis”, Journal of Proteomics (129) - Special Issue: Computational Proteomics: 51-55, 2015.
Diogo B Lima, Tatiani B de Lima, Tiago S Balbuena, Ana Gisele C. Neves-Ferreira, Valmir C Barbosa, Fabio C Gozzo, Paulo C Carvalho, “SIM-XL: a powerful and user-friendly tool for peptide cross-linking analysis”, Journal of Proteomics (129) - Special Issue: Computational Proteomics: 51-55, 2015.